A few months ago I launched a VHH binder design mini-competition. The itch I wanted to scratch was to see how well binder design tools do when run without hand-holding by the developers themselves—i.e., when run the way a typical user would.
There are more details in the original blogpost, but the gist was that the competitor submits a script to generate designs, and I run that script on a target.
If we had a "best script" for binder design, kind of like AlphaFold 3 is for folding, it would be hugely enabling for scientists.
I ended up allowing $100 of compute per design, which I thought was just on the edge of possibly producing a binder. It's also approximately the price of testing one design in the lab, which seems like a reasonable benchmark. The consensus from experts I talked to was that this would be insufficient to generate a binder. Turns out they were right! Nevertheless, here is the rundown.

Competitors
I convinced one person to enter this competition: Nick Boyd from Escalante Bio. Nick won the recent Adaptyv Nipah G competition using his own Mosaic protein design library (and it wasn't close!)
As you'd expect, Nick entered using a Mosaic script, similar to his Nipah G script, but adapted to generate a VHH instead of a mini-binder. While Mosaic is well validated for mini-binders, it has not really been tested for VHH designs, which are generally believed to be more difficult.
I entered using a BoltzGen script. My reasoning was that BoltzGen showed very strong results for VHH designs in their preprint, though they certainly used a lot more GPU hours than I did.
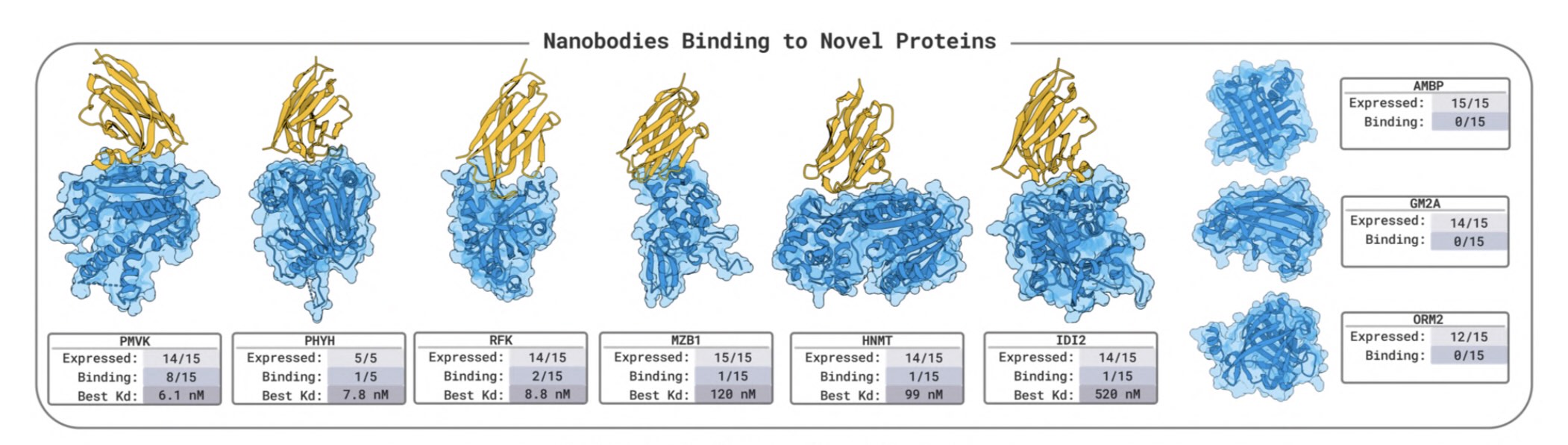 BoltzGen has arguably the strongest published VHH design results
BoltzGen has arguably the strongest published VHH design results
Results
I tested the designs against MBP, part of Adaptyv's BenchBB benchmark, which is a set of seven standardized targets designed to be used for benchmarking. If you elect to make the results public, as I did, you get a discount.
I posted the scripts and full results from Apaptyv on the competition github repo. The results should also appear on proteinbase.com in the near future. Of course, there is not much to see here, since none of the designs bound!
EasyMosaic
One complication of Mosaic compared to other tools like BindCraft, BoltzGen, or mBER is that Mosaic is a library, so the user is expected to define their own optimization parameters and loss function. For example, you could define a loss function as a weighted sum of ipTM, pLDDT, and distance to epitope. Different binder design problems might require a different balance of weights. This is a very powerful approach, and allows the user to tune Mosaic for different targets and use-cases, but it can be difficult to know where to start.
Part of the point of this competition was to see if Mosaic could be packaged into a user-friendly script. Since its success in the Nipah G competition, there has been quite a bit of interest in this.
With some advice from Nick on parameters, I made a web-based interface to mosaic called easymosaic. As with most of my stuff, it runs on modal and lets you run Mosaic with some reasonable default parameters for mini-binders or VHHs. The minibinder parameters should match the parameters used by Nick in the Nipah G competition.
Easymosaic is designed to do a decent job producing a binder without the need for parameter tuning. Your mileage will certainly vary a lot based on your target!
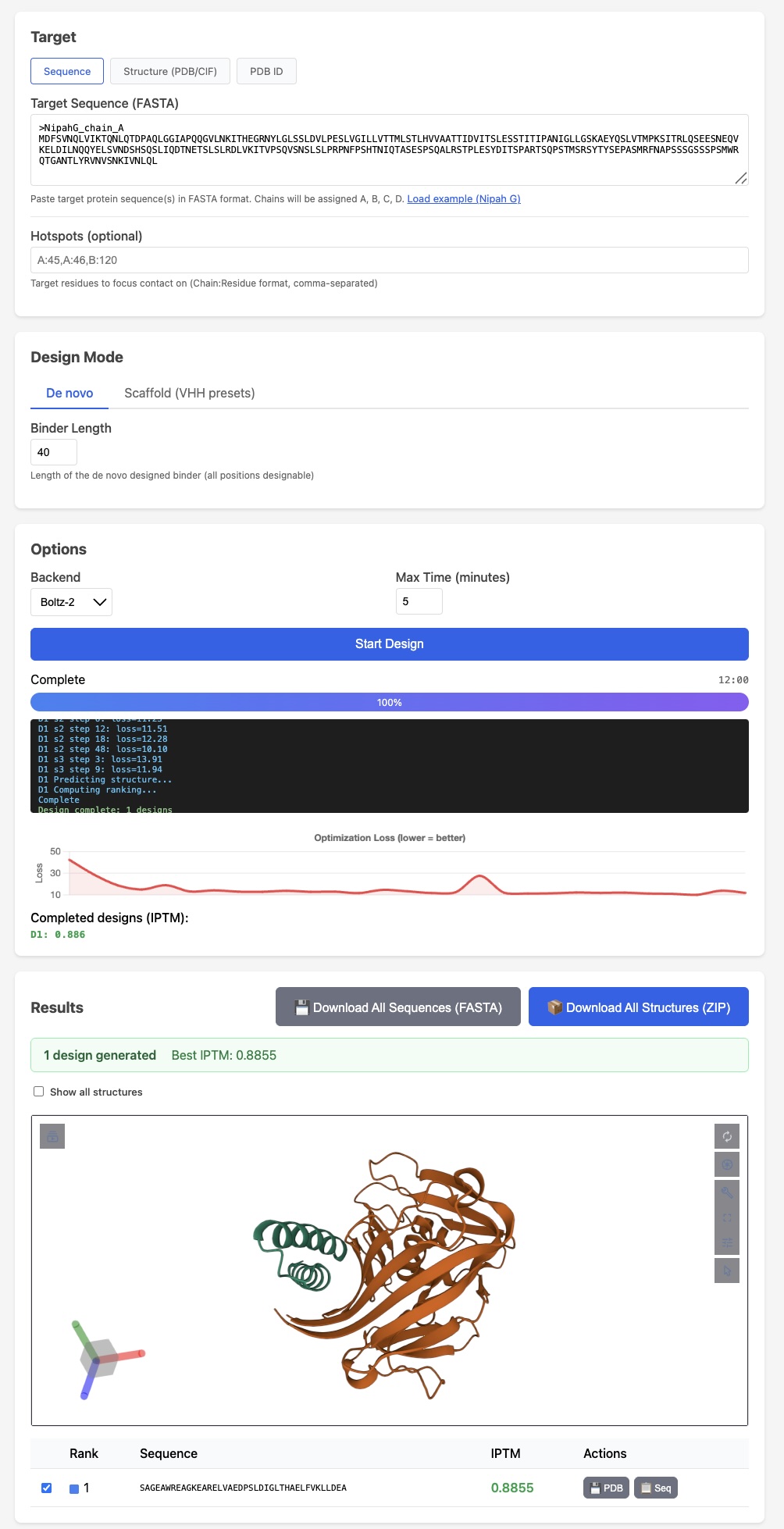 Like protein folding tools, easymosaic's interface has almost no options
Like protein folding tools, easymosaic's interface has almost no options
Mosaic-TUI
Nick's own Mosaic-TUI is a similar idea, but is more suitable for power users. It runs in the terminal, exposes all the relevant parameters, and has some nice features like the ability to use multiple GPUs.
Both easymosaic and Mosaic-TUI use B200 GPUs by default, so it is very easy to spend hundreds of dollars for a few good designs. Each design, before filtering out the bad ones, can cost $1 or more.

Sadly it's a bit too late to use either of these tools to enter the Adaptyv RBX1 competition but I'm sure there will be more competitions coming!
Hopefully, binder design tools will make some advances and I can try this again in a year or so, with a better chance of success. There are still plenty of things to try: combining the strengths of diffusion with hallucination; grounding designs in physics, etc.